mcPCR is designed to fill an important need in molecular pathology: PCR for methylation. Its arrival – announced at AGBT 2026 in February – could enable a new frontier in disease detection and monitoring, especially in cancer and many chronic diseases.
We asked Geoff Smith, Founder and CEO of Syndex Bio, to update us on the what, why, and how of mcPCR.

What’s the fundamental limitation of PCR in epigenetics?
Standard PCR copies and amplifies only the four-base DNA sequence. DNA methylation patterns, such as 5‑methylcytosine (5mC), are not picked up, and the epigenetic signal is effectively lost during amplification.
This means PCR alone cannot generate amplified material that still contains the original methylation patterns that are critical for detecting many diseases, including cancer.
How does mcPCR address this?
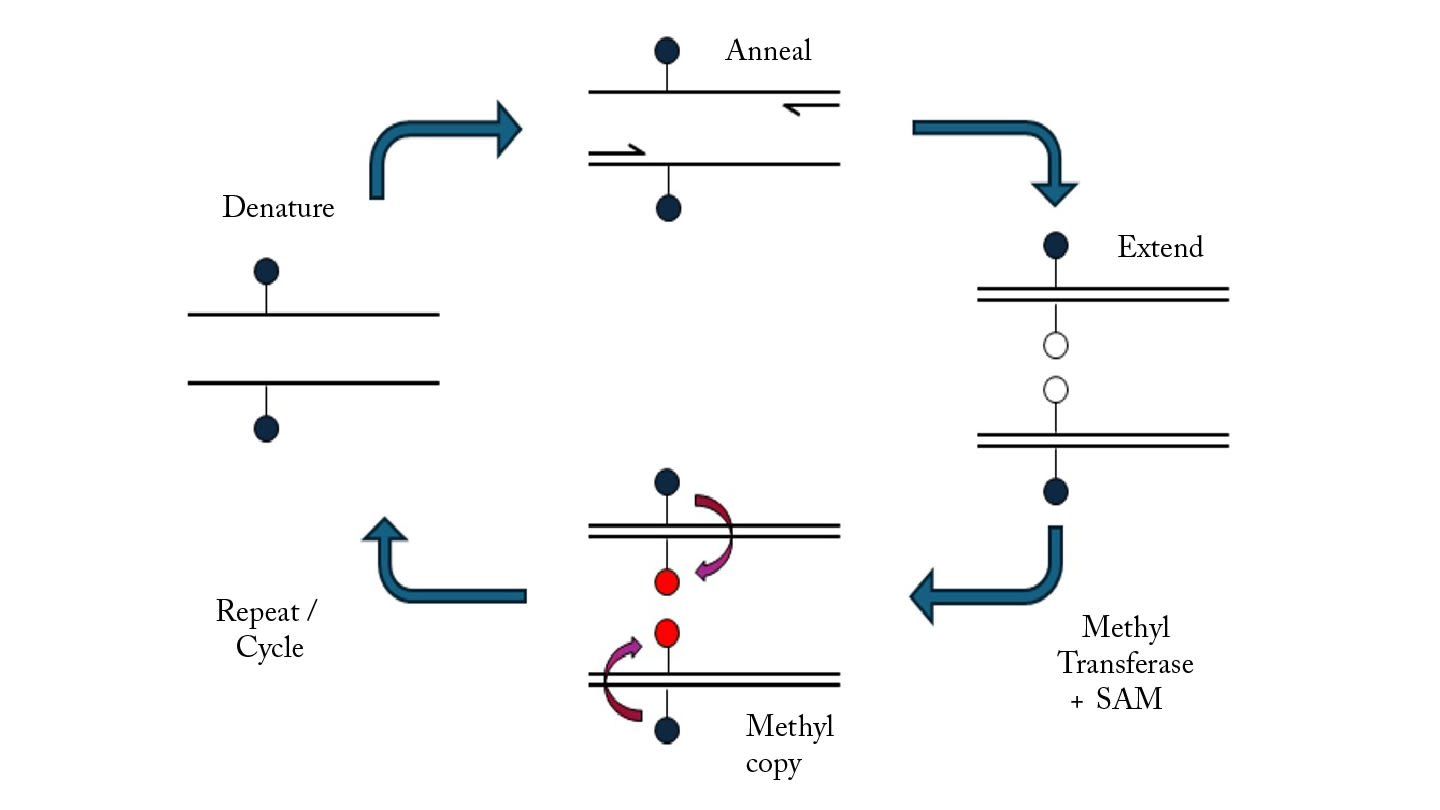
mcPCR couples high‑fidelity polymerase-mediated primer extension with a proprietary methyltransferase step that recopies 5mC onto newly synthesized strands after each cycle, converting hemi‑methylated intermediates into fully methylated products.
In doing so, it amplifies both the methylation patterns and the four-base genetic code, enabling simpler, faster, and more sensitive analysis of methylated DNA from limited clinical samples.
What challenges did you encounter in the development of mcPCR?
Technically, the challenge is to integrate enzymatic copying of methylation into a PCR-like cyclic workflow without sacrificing fidelity, yield, or compatibility with downstream sequencing and multi‑omic analyses.
There is also the translational challenge of engineering the platform for high-throughput, non-destructive workflows that work reliably on low-input, non-invasive, and small-sample biopsies, typical of early detection and recurrence monitoring settings.
How might mcPCR capability impact diagnostics?
By enabling amplification of methylation patterns, mcPCR could unlock more sensitive methylation-based assays for cancer, chronic disease, and residual disease monitoring, particularly when only tiny amounts of DNA are available.
This may broaden the use of methylation biomarkers beyond specialized sequencing-only pipelines, supporting earlier and more accurate detection from blood and other minimally invasive samples.
How might it impact patients?
If methylation can be read out more sensitively from non-invasive and small biopsies, clinicians could detect disease earlier, monitor response and recurrence more precisely, and potentially reduce the need for more invasive tissue procedures.
For patients, that could translate into earlier intervention, better tailoring of therapy, and more comfortable longitudinal monitoring over the course of treatment and survivorship.




